Credit: Martin Christoph Konnerth
Is is possible to develop antibiotics with the help of bacteria from in our own body? Perhaps. This week’s issue of Nature presented a discovery of how a bacteria found in our noses seems to be able to fight other bacteria, including the resistant MRSA bacteria.
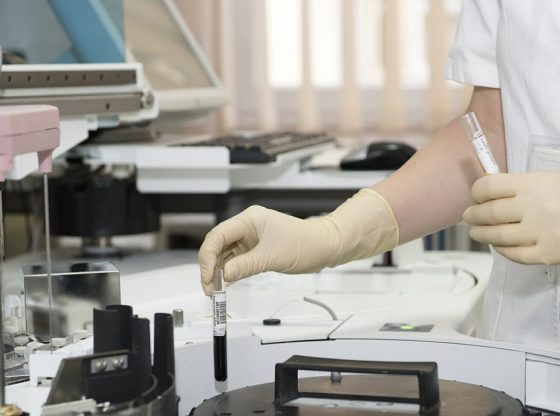
Methicillin-resistant Staphylococcus aureus (MRSA) is a bacterium responsible for several difficult-to-treat infections in humans. It can result in severe infections of the blood and heart.
More Antibiotics = Faster Evolution
The problem of resistant bacteria is surmounting, with more antibiotics becoming ineffective. Resistance arises through one of three ways: natural resistance in certain types of bacteria; genetic mutation; or by one species acquiring resistance from another.
Over usage of antibiotics in treating humans is greatly worsening the problem, as antibiotics are heavily used in many parts of the world at times when a smaller dose or none at all would have been sufficient. Recently the Indonesian health minister called on doctors from over prescribing antibiotics – where according to research by the Antimicrobial Resistance Control Committee (KPRA) at the Health Ministry – more than 50 percent of antibiotic prescriptions from doctors are unnecessary.
The Netherlands has the lowest rate of antibiotic prescribing in the OECD, at a rate of 11.4 defined daily doses (DDD) per 1,000 people per day in 2011. Germany and Sweden also have lower prescribing rates. By contrast, Greece, France, and Belgium have high prescribing rates of more than 28 DDD.
More problematic however is the use of antibiotics for purposes in the husbandry of livestock, this includes not only the treatment or prophylaxis of infection but also the use of subtherapeutic doses in animal feed and/or water to promote growth and improve feed efficiency in intensive animal farming.
That is, healthy animals are receiving antibiotics as a precaution. Since the industry is characterized by animals in very close proximity to each other and often their excrement, it is an environment where bacteria should thrive, but instead of keeping the environment clean, the animals are often given antibiotics instead, giving rise to mutating resistant bacteria.
Friendly Nose Bacteria
The hunt for new antibiotics is on and this new paper presents encouraging findings of perhaps harnessing bacteria found in ourselves to be used as a future antibiotic.
The researchers looked into the nose of more than 180 patients. Approximately ten percent of them had a kind type of staph bacteria, one called Staphylococcus lugdunensis.
Almost everyone who had this variant lacked the most dangerous variant of Staphylococcus aureus, which can become resistant to common antibiotics (MRSA).
This symbiotic bacteria as it seems produces a variant of antibiotic substance that scientists have dubbed lugdunin. The researchers further examined this substance via experiments on mice injected with lugdunin and it showed promising results – able to cure skin infections caused by the more problematic aureus variant.
“It is still unclear what benefit this newly discovered antibiotic can do to cure more serious infections caused by MRSA”, study author Diarmaid Hughes said.
He concludes that it is certainly an exciting approach to looking for antibiotics in our own bodies. But adds that there are still many more bacteria to explore in many other different environments, for example, in the Earth’ soil – if we are to combat the problem of antibiotic resistance.
_____________
Zipperer Alexander et al. Human commensals producing a novel antibiotic impair the pathogen colonization. Nature, 2016. DOI: 10.1038 / nature18634.
__________________________